2. 济南大学土木建筑学院,济南 250022;
3. 许家崖水库管理处,山东临沂 273400
2. School of Civil Engineering and Architecture, University of Jinan, Jinan 250022, China;
3. Xujiaya Reservoir Management Office, LinYi, Shandong 273400, China
中国一直积极参与国际气候变化治理及全球减排进程,在《巴黎协定》上提出我国碳减排目标:2020年碳排放强度相比2005年降低40%~45%,2030年碳排放强度相比2005年下降60%~65%[1]。因此,我们要积极采取措施加快碳减排进程,大力减少碳排放总量。湿地是大气中CH4最大的天然排放源[2]。近年来,CH4浓度以每年约6×10–3cm3 m–3的速度剧烈上升[3],2018年1—10月全球平均温度较工业化前高0.98±0.1℃,是有记录以来第四最温暖的年份,仅次于发生了超级厄尔尼诺的2015年、2016年和2017年,CH4浓度达到了1 859(±10)×10–3cm3 m–3,是工业化前水平的257%[4]。创纪录的温室气体浓度将全球温度推向越来越危险的水平,不仅影响到全球气候变化,而且气候变化的反馈作用会破坏生态系统的稳定性,从而进一步推动变暖。
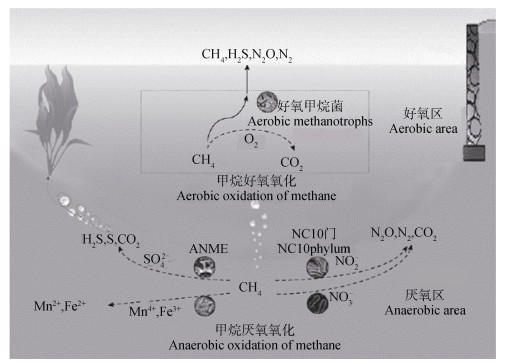
CH4的氧化分为两大类型(图 1):一类是有氧条件下由微生物利用O2作为电子受体将CH4氧化;另一类是在厌氧条件下由微生物利用除O2外的其他电子受体(如
|
注:ANME,甲烷厌氧氧化古菌。 Note: ANME, Anaerobic methanotrophic archaea. 图 1 CH4氧化原理[5] Fig. 1 Methane oxidation principle[5] |
目前,根据电子受体的不同,AOM途径主要为硫酸盐还原型甲烷厌氧氧化(SAMO),反硝化型甲烷厌氧氧化(DAMO),以Fe3+、Mn4+作为电子受体的甲烷厌氧氧化(Metal-AOM)以及种间直接电
子转移(DIET)。
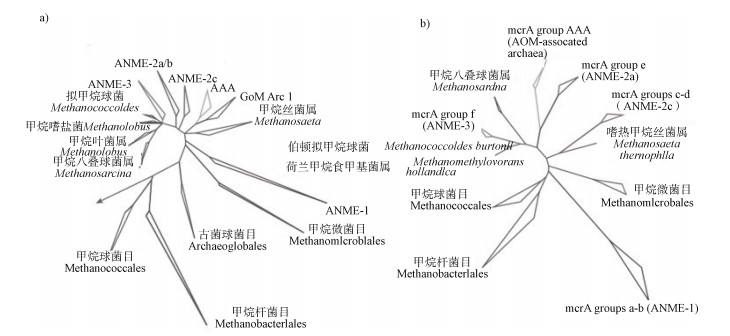
1.1 硫酸盐还原型甲烷厌氧氧化(SAMO)SAMO过程的微生物包含甲烷厌氧氧化古菌(ANME)和硫酸盐还原细菌(SRB)。这两类微生物在空间上紧密耦合联系[12],通常以聚集体的形式存在[13]。研究者们基于16S rRNA和改良胞嘧啶限制蛋白A(mcrA)基因多态性,将ANME划分为3类:ANME-1、ANME-2、ANME-3,均属于广古菌门,它们的亲缘关系如图 2a)[14]:ANME1与产甲烷微菌目(Methanomicrobiale)有较远的亲缘关系;ANME-2属于产甲烷八叠球菌目(Methanosarcinales),包含4个亚簇:ANME-2a、ANME-2b、ANME-2c、ANME-2d(又叫AAA);而ANME-3与甲烷拟球菌属(Methanococcoides)亲缘关系较近。这3类ANME的序列相似度仅有75%~92%,但生理特性相似,表明不同的ANME群落偏好不同的生态环境。ANME- mcrA基因的系统发育分析显示,ANME-1—3的亲缘关系十分疏远(图 2b)),与16S rRNA结果相符合。
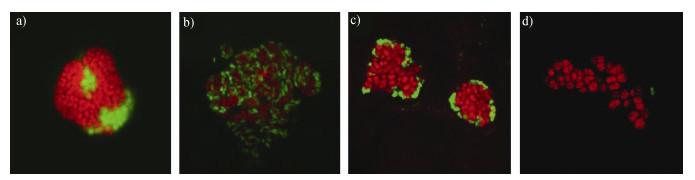
SRB种类丰富,在SAMO过程中主要包括脱硫八叠球菌属(Desulfosarcina)、脱硫球菌属(Desulfococcus)和脱硫叶菌属(Desulfobuibus)。脱硫八叠球菌属(Desulfosarcina:DSS)和脱硫球菌属(Desulfococcus)主要吸附在ANME-1、ANME-2上,形成细胞絮凝体[15-16];脱硫叶菌属一般与ANME-3共存发生作用(图 3a))。ANME-2与SRB组成互养共生体,其内部核心由ANME-2充当,而其外部被SRB构成的壳层部分或全部包围(图 3b),图 3c))[17]。随着进一步的研究发现,SRB与ANME的共存形式呈现多样化,如在Eel河流域沉积物中发现了脱硫叶菌属与ANME-2c共存现象,以及脱硫八叠球菌属和脱硫球菌属与ANME-2c共存的现象[18];更多的证据表明与ANME共生的细菌可能不仅限于SRB,而是具有较高的多样性[19],如仅由ANME-2细胞形成的单一类型微生物的共生体(图 3d))[20-21];在某些环境发现α-变形菌纲的Spbingomonas spp.和β-变形菌纲的Burkbolderia spp.也会与ANME-2c关联生长[18]。
|
图 3 通过荧光原位杂交(FISH)显示的不同类型甲烷厌氧氧化(AOM)古菌团聚体的荧光显微照片(a)ANME-3(红色)与脱硫叶菌(绿色)团聚体;b)渗漏沉积物中观察到ANME-2a(红色)和脱硫八叠球菌属(DSS)(绿色)细胞的混合型聚生体;c)渗漏沉积物中的ANME-2c(红色)和DSS(绿色)细胞的壳型聚生体;d)渗漏沉积物中单个ANME-2c细胞[14].) Fig. 3 Epifluorescence micrographs of different species of anaerobic methanotrophic archaea(ANME)aggregates visualized by fluorescence in situ hybridization(FISH)(a)ANME-3(red)and Desulfobulbus(green)consortia; b)Mixed-type consortia of ANME-2a(red)and Desulfosarcina(DSS)(green)cellso bserved in sediments; c)Shell-type consortia of ANME-2c(red)and DSS(green)cells in sediments; d)single ANME-2c cells observed in sediments[14].) |
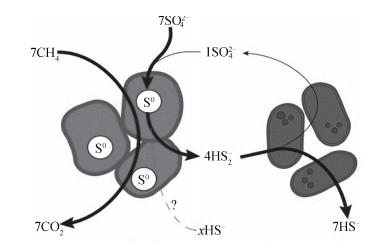
当前,SAMO理论反应机理主要有四种(表 1):反向产甲烷理论、乙酸生成理论、甲基化理论和以S0作为中间产物(图 4)的SAMO机理[28-39]。其中,反向产甲烷理论模型研究得最深入[28-34]。
|
|
表 1 硫酸盐还原型甲烷厌氧氧化四种理论模型 Table 1 Introduction of four theoretical models of sulfate reducing anaerobic oxidation of methane(SAMO) |
|
图 4 以S0作为中间产物的甲烷厌氧氧化[28] Fig. 4 Anaerobic oxidation of methane with S0 as the intermediate product28] |
乙酸生成理论是对反向产甲烷理论的补充与发展,它解释了H2在AOM过程中并非起中间体作用,H2浓度与AOM作用之间的影响关系。以S单质作为中间产物的SAMO机制被认作是一个全新的途径[30],该过程由ANME的一个分支(ANME-2)在无SRB参与下,以S0为中间载体完成了AOM过程中的硫酸盐还原,从一个全新的角度定义ANME和SRB的功能,开拓了一个新的研究方向。
1.2 反硝化型甲烷厌氧氧化(DAMO)参与DAMO途径的微生物是一类新的微生物Candidatus Methylomirabilis oxyfera(M. oxyfera)[17],隶属于新发现的细菌门—NC10门,且所有与反硝化过程耦合的AOM富集培养中均含有一定量(30%~80%)的NC10门细菌;其他少数古菌为ANME(属于甲烷八叠球菌目)的AAA分支的细菌。NC10门细菌细胞呈细杆状、杆状,以单细胞或团聚体的形式存在。根据NC10门设计的特异引物,将基因库中的序列比对之后发现,NC10门细菌包括a、b、c及d四个类群。然而,目前所富集的细菌均归属于a类群,说明a类群是参与DAMO的主要功能群,而NC10门中其他种类的细菌是否具有甲烷厌氧氧化的能力尚未确定。
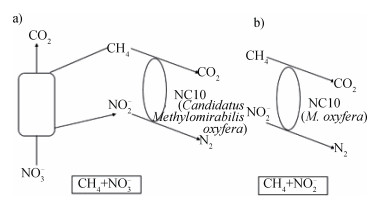
目前针对DAMO机理的研究分为NC10门细菌单独催化DAMO的催化机理和古菌协同催化机理两个研究方向(图 5)[31]。
|
图 5 |
(1)NC10门细菌催化DAMO机理2006年,Raghoebarsing等[32]通过全基因测序推测出DAMO的机理:基于M. oxyfera本身缺少编码NO还原酶的基因却能够自己生成O2的实验结果,推测它的细胞内存在某种能够歧化NO生成O2的未知酶,将产生的一部分(3/4)O2用于CH4的氧化反应;2011年Wu等[33]提出了M. oxyfera细菌的分解代谢和能量代谢途径:在亚硝酸还原酶的作用下,
(2)古菌参与DAMO的催化机理目前,对于古菌参与DAMO途径的研究较少。在Raghoebarsing[32]、Hu等[35]的文章中提到DAMO的富集培养物中存在着古菌和细菌的混合物,且均具有消耗
DAMO的研究进展如表 2所示,这是一种近年新发现的AOM途径,在微生物学、生态学和环境工程领域有很大的研究价值和前景,发现至今受到众多学者的关注。研究者致力于分析DAMO功能微生物、功能微生物的内在机理及其富集的影响因素和DAMO的实际应用。因DAMO中主要功能微生物M. oxyfera富集培养缓慢,时间长达数周甚至数月,反硝化速率较低,目前工艺研究较少,主要应用DAMO脱氮工艺[36]:利用DAMO功能菌与Anammox功能菌共同作用进行脱氮,CH4作为反硝化过程中的电子供体,将
|
|
表 2 反硝化型甲烷厌氧氧化的相关研究进展 Table 2 Progresses of researches on denitrifying anaerobic oxidation of methane |
Metal-AOM反应涉及固、液、气相,反应速率低且机理复杂。早期研究中,Beal等[49]怀疑ANME-1、ANME-3或拟甲烷球菌可能与甲烷氧化有关,金属还原细菌(如某些种类的拟杆菌、脱硫单胞菌、酸杆菌)可能作为它们的伙伴,对金属还原起着至关重要的作用。根据对台湾东部陆地泥火山中AOM和微生物群落的环境调查,研究人员认为ANME-1~ANME-3和脱硫单胞菌属可能与Metal-AOM有关[50]。在实验室培养微生物研究Metal-AOM菌群的报道中提到属于ANME的ANME-2亚类ANME-2a、ANME-2c、ANME-2d(AAA)均可参与Metal-AOM。2016年,Scheller等[51]、Fu等[52]及Ettwig等[53]的三项独立研究表明,一些菌株的ANME-2无论是否有细菌伴侣,均能够结合CH4进行金属还原,且ANME-2d能够在不同的环境条件下利用不同的氧化剂作为电子受体。
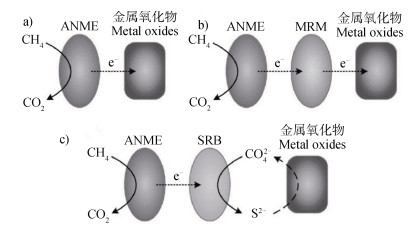
目前主要的机理有三种:第一种通过几种微生物(ANME-2a、ANME-2c、ANME-2d),负责Metal- AOM的整个过程,ANME氧化CH4是将电子直接转移至可溶性金属离子、复合物或者固体金属氧化物(图 6a))。第二种氧化机理与SAMO一样,ANME与金属还原微生物(MRM)合作进行金属的厌氧氧化,相关的MRM接受来自ANME的电子以分解金属氧化物(图 6b))。值得注意的是,在SAMO机理中发生的“隐硫循环”(在硫化物存在条件下形成S0,通过S0的歧化反应生成
|
注:MRM:金属还原微生物。 Note: MRM: Metal reducing microorganism. 图 6 不同机制下微生物介导的金属甲烷厌氧氧化[54] Fig. 6 Different mechanisms on microbe-mediated metal anaerobic oxidation of methane [54] |
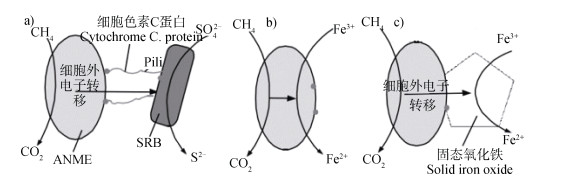
种间直接电子转移(DIET)是在Milucka等[65]和Nauhaus等[12]研究基础上推断出的一种新的AOM途径。他们通过对海洋中的ANME菌和SRB菌还原硫酸盐的实验猜测ANME与SRB之间的电子转移体系不需要通过常规的中间产物(间氢、醋酸或者甲酸等)作为电子转移载体,而是一种更直接的方式——DIET。DIET的前身是胞外电子转移机制(DEET)[66],最早发现于地杆菌的共培养中;随后,Summers等[67]又在金属还原性土壤杆菌和硫还原性土壤杆菌共培养过程中发现两种菌体形成了一种导电的聚集体,聚集体DNA测序结果表明,在缺乏种间氢转移的突变体中,团聚体的形成较快,证明种间电子转移是直接的;此外,产甲烷过程中也发生DIET(图 7)[68]。随着近年来稳定同位素示踪、宏基因组、理论模型等先进技术的发展和AOM上的应用,2015年Wegener等[69]用ANME-1古细菌和SRB(HotSeep-1)在60℃下测试直接种间电子交换的假设,观察到在嗜热AOM(TAOM)条件下,古菌和细菌均过度表达细胞色素生成的基因,并形成类似于种间电子连接的纳米线结构,这种结构负责共生菌群之间的种间电子转移;McGlynn等[70]通过单细胞色素的检测以及聚集体细胞之间基质的氧化还原依赖性染色实验,为直接电子转移的共同耦合提供了证据;Scheller等[51]使用速率测量和单细胞稳定同位素探测相结合的方法证明SAMO过程无硫酸盐时,AOM反应速率并未衰减,正是通过DIET这种方式进行电子交换。
|
|
表 3 金属-甲烷厌氧氧化研究 Table 3 Studies on metal anaerobic oxidation of methane |
|
图 7 种间直接电子转移机制下甲烷厌氧氧化(AOM)电子转移方式[68] Fig. 7 Electron transfer in anaerobic oxidation of methane controlled by direct interspecies electron transfer(DIET)[68] |
传统的微生物多样性研究方法是将微生物从环境中分离,然后在实验室对其进行分离培养得到微生物的单菌株。由于微生物种类繁多,实验室富集培养微生物的仅有极少数,造成了研究上的困难。而现代微生物多样性研究是基于Illumina HiSeq测序平台,利用双末端测序的方法,构建小片段文库在编码核糖体RNA的核酸序列保守区进行测序。通过对序列拼接过滤,操作分类单元(Operational taxonomic units,OTU)聚类,并进行物种注释及丰度分析,揭示样品的物种构成;进一步进行α多样性分析、β多样性分析和显著物种差异分析等,挖掘样品之间的差异。
2.1 现代检测方法分类及优点现代分子生物学技术能够揭示DNA序列遗传多态性,用于推断系统发育关系,并通过数据库比较鉴定未知微生物,其中基于16S rRNA的分子生物学技术已被广泛应用(表 4),例如变性梯度凝胶电泳(DGGE)、温度梯度凝胶电泳(TGGE)、末端限制性片段长度多态性分析(T-RFLP)等可依据独特核酸物质的分离,提供微生物多样性的轮廓;宏基因组学由Handelsman等[71]1998年提出,对生境中所有微生物的总基因组DNA进行研究,可避开传统可培养方法的局限性,提高新微生物的发现效率,结合宏转录组学及新一代质谱技术催生下的宏蛋白质组学,可系统研究微生物群落及其功能,全面分析其组成、发展规律、进化和功能预测,认识微生物群落中物种多样性、基因潜力以及基因表达与调控,已广泛应用于海洋湖泊、深海热泉、人体肠道、牛瘤胃生境、森林土壤与堆肥生境等环境中微生物群落的结构和功能研究。
|
|
表 4 几种分子生物学技术的应用情况 Table 4 Application of several molecular biological methods |
分子生物学技术的发展和进步促进了环境微生物的研究,但不同方法有其局限性,如PCR扩增以优势种群为主,不利于检测丰度较低的稀有微生物种群;高通量测序得到的序列难以组成完整的基因,且很多基因的功能尚不清楚,很难获得由序列到功能的准确图谱;DGGE中受影响的条带可能含有多个物种,检测结果需要结合克隆库来验证;454测序中对一条序列的同一个碱基的重复测量准确度低;Illumina测序在测定GC含量(鸟嘌呤和胞嘧啶的比率)高的序列区域时容易产生偏差;FISH样品的荧光会影响探针的荧光信号,导致微生物细胞的同位素组成受到样品制备过程的干扰。
3 展望本文综述了四种不同电子受体的湿地AOM途径以及参与反应的微生物菌群,探讨了环境因子对湿地AOM的影响机制,列举了几种研究微生物群落结构特征、数量分布特征、活性和相对作用强度的分子生物学技术,为从原理上探明AOM途径的发生机理和微生物在功能、生态和进化上的关系提供更多的科学依据,在湿地温室气体排放领域取得新突破。关于湿地AOM途径的未来研究方向可从以下几个方面进行。
首先,微生物的富集和纯培养是深入研究AOM的基础,但由于功能微生物的培养过程非常缓慢,目前针对AOM各过程机理的研究多数仍停留在理论阶段,多种理论间互有冲突,未有充足的实验证据支撑,其中对于Metal-AOM途径以及DIET途径的研究更是处于起步阶段。今后的研究需要进一步探索各途径微生物菌群富集培养的适宜条件,减少各过程之间的相互影响,提高反应稳定性。
其次,由于河口湿地受到淡水与海水入侵的双重作用,以及人类活动对湿地碳源输入的影响,湿地AOM过程的环境影响因子(温度、盐度、植被类型、pH等)和电子受体(
最后,AOM途径应用于实际环境工程。例如可利用DAMO处理目前亟待解决的含氮废水,将CH4作为替代碳源用于传统的反硝化工艺,并研发新型的脱氮除碳工艺,具有明显的经济效益和环境生态效益,可进一步研发具体的工艺流程。
| [1] |
Wang Y, Cheng Y, Yang G C, et al. Provincial decomposition of China's carbon emission rights under the constraint of 2020 and 2030 carbon intensity targets (In Chinese)[J]. China Environmental Science, 2018, 38(8): 3180-3188. DOI:10.3969/j.issn.1000-6923.2018.08.049 [王勇, 程瑜, 杨光春, 等. 2020和2030年碳强度目标约束下中国碳排放权的省区分解[J]. 中国环境科学, 2018, 38(8): 3180-3188.]
(  0) 0) |
| [2] |
Kim S Y, Veraart A J, Meima-Franke M, et al. Combined effects of carbon, nitrogen and phosphorus on CH4, production and denitrification in wetland sediments[J]. Geoderma, 2015, 259/260: 354-361. DOI:10.1016/j.geoderma.2015.03.015
(  0) 0) |
| [3] |
Nisbet E G, Dlugokencky E J, Bousquet P. Methane on the rise-again[J]. Science, 2014, 343(6170): 493-495. DOI:10.1126/science.1247828
(  0) 0) |
| [4] |
Petteri T, António G. The State of the Global Climate in 2018. World Meteorological Organization, https://public.wmo.int, 2018-12.
(  0) 0) |
| [5] |
Thauer R. Functionalization of methane in anaerobic microorganisms[J]. Angewandte Chemie-International Edition, 2010, 49(38): 6712-6713. DOI:10.1002/anie.201002967
(  0) 0) |
| [6] |
Martens C S, Berner R A. Methane production in the interstitial waters of sulfate-depleted marine sediments[J]. Science, 1974, 185(4157): 1167-1169. DOI:10.1126/science.185.4157.1167
(  0) 0) |
| [7] |
Ding J, Lu Y Z, Fu L, et al. Decoupling of DAMO archaea from DAMO bacteria in a methane-driven microbial fuel cell[J]. Water Research, 2017, 110: 112-119. DOI:10.1016/j.watres.2016.12.006
(  0) 0) |
| [8] |
Segarra K E A, Schubotz F, Samarkin V, et al. High rates of anaerobic methane oxidation in freshwater wetlands reduce potential atmospheric methane emissions[J]. Nature Communications, 2015, 6: Article number 7477. DOI:10.1038/ncomms8477
(  0) 0) |
| [9] |
Radajewski S, Ineson P, Parekh N R, et al. Stable-isotope probing as a tool microbial ecology[J]. Nature, 2000, 403(6770): 646-649. DOI:10.1038/35001054
(  0) 0) |
| [10] |
Ji H F, Wang Y. Advances in molecular approach applications in microbial ecology studies (In Chinese)[J]. Acta Ecologica Sinica, 2016, 36(24): 8234-8243. [姬洪飞, 王颖. 分子生物学方法在环境微生物生态学中的应用研究进展[J]. 生态学报, 2016, 36(24): 8234-8243.]
(  0) 0) |
| [11] |
Liu L Y, Cui H F, Tian G. Application of next generation sequencing technology in macrogenomics (In Chinese)[J]. Chinese Medicinal Biotechnology, 2013, 8(3): 196-200. DOI:10.3969/cmba.j.issn.1673-713X.2013.03.008 [刘莉扬, 崔鸿飞, 田埂. 高通量测序技术在宏基因组学中的应用[J]. 中国医药生物技术, 2013, 8(3): 196-200.]
(  0) 0) |
| [12] |
Nauhaus K, Treude T, Boetius A, et al. Environmental regulation of the anaerobic oxidation of methane:A comparison of ANME-Ⅰ and ANME-Ⅱ communities[J]. Environmental Microbiology, 2005, 7(1): 98-106.
(  0) 0) |
| [13] |
Alperin M J, Hoehler T M. Anaerobic methane oxidation by archaea/sulfate-reducing bacteria aggregates:1. Thermodynamic and physical constraints[J]. American Journal of Science, 2009, 309(10): 869-957. DOI:10.2475/10.2009.01
(  0) 0) |
| [14] |
Knittel K, Boetius A. Anaerobic oxidation of methane:Progress with an unknown process[J]. Annual Review of Microbiology, 2009, 63(1): 311-334.
(  0) 0) |
| [15] |
Niemann H, Lösekann T, de Beer D, et al. Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink[J]. Nature, 2006, 443(7113): 854-858. DOI:10.1038/nature05227
(  0) 0) |
| [16] |
Losekann T, Knittel K, Nadalig T, et al. Diversity and abundance of aerobic and anaerobic methane oxidizers at the Haakon Mosby mud volcano, Barents Sea[J]. Applied and Environmental Microbiology, 2007, 73(10): 3348-3362. DOI:10.1128/AEM.00016-07
(  0) 0) |
| [17] |
Knittel K, Losekann T, Boetius A, et al. Diversity and distribution of methanotrophic archaea at cold seeps[J]. Applied and Environmental Microbiology, 2005, 71(1): 467-479.
(  0) 0) |
| [18] |
Pernthaler A, Dekas A E, Brown C T, et al. Diverse syntrophic partnerships from deep-Sea methane vents revealed by direct cell capture and metagenomics[J]. Proceedings of the National Academy of Sciences of the United States of America, 2008, 105(19): 7052-7057. DOI:10.1073/pnas.0711303105
(  0) 0) |
| [19] |
Sun Z L, He Y J, Li J, et al. Progress and environmental effect in seafloor anaerobic oxidation of methane (In Chinese)[J]. Advances in Earth Science, 2012, 27(11): 1262-1273. [孙治雷, 何拥军, 李军, 等. 海洋环境中甲烷厌氧氧化机理及环境效应[J]. 地球科学进展, 2012, 27(11): 1262-1273.]
(  0) 0) |
| [20] |
Orphan V J, House C H, Hinrichs K U, et al. Multiple archaeal groups mediate methane oxidation in anoxic cold seep sediments[J]. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(11): 7663-7668. DOI:10.1073/pnas.072210299
(  0) 0) |
| [21] |
Treude T, Orphan V, Knittel K, et al. Consumption of methane and CO2 by methanotrophic microbial mats from gas seeps of the anoxic Black Sea[J]. Applied and Environmental Microbiology, 2007, 73(7): 2271-2283. DOI:10.1128/AEM.02685-06
(  0) 0) |
| [22] |
Krüger M, Meyerdierks A, Glöckner F O, et al. A conspicuous nickel protein in microbial mats that oxidize methane anaerobically[J]. Nature, 2003, 426(6968): 878-881. DOI:10.1038/nature02207
(  0) 0) |
| [23] |
Hoehler T M, Alperin M J, Albert D B, et al. Field and laboratory studies of methane oxidation in an anoxic marine sediment:Evidence for a methanogen-sulfate reducer consortium[J]. Global Biogeochemical Cycles, 1994, 8(4): 451-463. DOI:10.1029/94GB01800
(  0) 0) |
| [24] |
Scheller S, Goenrich M, Boecher R, et al. The key nickel enzyme of methanogenesis catalyses the anaerobic oxidation of methane[J]. Nature, 2010, 465(7298): 606-608. DOI:10.1038/nature09015
(  0) 0) |
| [25] |
Valentine D L, Reeburgh W S. New perspectives on anaerobic methane oxidation[J]. Environmental Microbiology, 2000, 2(5): 477-484. DOI:10.1046/j.1462-2920.2000.00135.x
(  0) 0) |
| [26] |
Moran J J, Beal E J, Vrentas J M, et al. Methyl sulfides as intermediates in the anaerobic oxidation of methane[J]. Environmental Microbiology, 2008, 10(1): 162-173.
(  0) 0) |
| [27] |
Lessner D J, Li L, Li Q, et al. An unconventional pathway for reduction of CO2 to methane in CO-grown Methanosarcina acetivorans revealed by proteomics[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103(47): 17921-17926. DOI:10.1073/pnas.0608833103
(  0) 0) |
| [28] |
Milucka J, Ferdelman T G, Polerecky L, et al. Zero-valent sulphur is a key intermediate in marine methane oxidation[J]. Nature, 2012, 491(7425): 541-546. DOI:10.1038/nature11656
(  0) 0) |
| [29] |
Wankel S D, Adams M M, Johnston D T, et al. Anaerobic methane oxidation in metalliferous hydrothermal sediments:Influence on carbon flux and decoupling from sulfate reduction[J]. Environmental Microbiology, 2012, 14(10): 2726-2740. DOI:10.1111/j.1462-2920.2012.02825.x
(  0) 0) |
| [30] |
Schreiber L, Holler T, Knittel K, et al. Identification of the dominant sulfate-reducing bacterial partner of anaerobic methanotrophs of the ANME-2 clade[J]. Environmental Microbiology, 2010, 12: 2327-2340.
(  0) 0) |
| [31] |
Haroon M F, Hu S H, Shi Y, et al. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage[J]. Nature, 2013, 500(7464): 567-570. DOI:10.1038/nature12375
(  0) 0) |
| [32] |
Raghoebarsing A A, Pol A, van de Pas-Schoonen K T, et al. A microbial consortium couples anaerobic methane oxidation to denitrification[J]. Nature, 2006, 440(7086): 918-921. DOI:10.1038/nature04617
(  0) 0) |
| [33] |
Wu M L, Ettwig K F, Jetten M S M, et al. A new intra-aerobic metabolism in the nitrite-dependent anaerobic methane-oxidizing bacterium Candidatus 'Methylomirabilis oxyfera'[J]. Biochemical Society Transactions, 2011, 39(1): 243-248.
(  0) 0) |
| [34] |
Wu M L, Alen T A, Donselaar E G, et al. Co-localization of particulate methane monooxygenase and cd1 nitrite reductase in the denitrifying methanotroph 'Candidatus Methylomirabilis oxyfera'[J]. FEMS Microbiology Letters, 2012, 334(1): 49-56.
(  0) 0) |
| [35] |
Hu S H, Zeng R J, Keller J, et al. Effect of nitrate and nitrite on the selection of microorganisms in the denitrifying anaerobic methane oxidation process[J]. Environmental Microbiology Reports, 2011, 3(3): 315-319.
(  0) 0) |
| [36] |
Shi Y, Hu S H, Lou J Q, et al. Nitrogen removal from wastewater by coupling anammox and methane- dependent denitrification in a membrane biofilm reactor[J]. Environmental Science & Technology, 2013, 47(20): 11577-11583.
(  0) 0) |
| [37] |
He Z F, Geng S, Pan Y W, et al. Improvement of the trace metal composition of medium for nitrite-dependent anaerobic methane oxidation bacteria:Iron(Ⅱ)and copper(Ⅱ)make a difference[J]. Water Research, 2015, 85: 235-243.
(  0) 0) |
| [38] |
Wang J S, Zhang Z J, Wang X J, et al. A review on artificial enrichment and application of denitrifying anaerobic methane oxidizing microorganisms (In Chinese)[J]. Technology of Water Treatment, 2017, 43(12): 8-13. [王劲松, 张召基, 王晓君, 等. 反硝化型甲烷厌氧氧化微生物人工富集与应用研究进展[J]. 水处理技术, 2017, 43(12): 8-13.]
(  0) 0) |
| [39] |
Yang M X, Guo Q W, Tong T L, et al. Vegetation type and layer depth influence nitrite-dependent methane-oxidizing bacteria in constructed wetland[J]. Archives of Microbiology, 2017, 199(3): 505-511.
(  0) 0) |
| [40] |
Shen L, Wu H, Gao Z, et al. Presence of diverse Candidatus Methylomirabilis oxyfera-like bacteria of NC10 Phylum in agricultural soils[J]. Journal of Applied Microbiology, 2016, 120(6): 1552-1560.
(  0) 0) |
| [41] |
Shen L D, Wu H S, Gao Z Q, et al. Comparison of community structures of Candidatus Methylomirabilis oxyfera-like bacteria of NC10 Phylum in different freshwater habitats[J]. Scientific Reports, 2016, 6: 25647.
(  0) 0) |
| [42] |
Padilla C C, Bristow L A, Sarode N, et al. NC10 bacteria in marine oxygen minimum zones[J]. The ISME Journal, 2016, 10(8): 2067-2071.
(  0) 0) |
| [43] |
He Z F, Geng S, Shen L D, et al. The short- and long-term effects of environmental conditions on anaerobic methane oxidation coupled to nitrite reduction[J]. Water Research, 2015, 68: 554-562.
(  0) 0) |
| [44] |
Hu B L, Shen L D, Lian X, et al. Evidence for nitrite-dependent anaerobic methane oxidation as a previously overlooked microbial methane sink in wetlands[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(12): 4495-4500.
(  0) 0) |
| [45] |
Zhu G B, Zhou L L, Wang Y, et al. Biogeographical distribution of denitrifying anaerobic methane oxidizing bacteria in Chinese wetland ecosystems[J]. Environmental Microbiology Reports, 2015, 7(1): 128-138.
(  0) 0) |
| [46] |
Zhu J, Wang Q, Yuan M D, et al. Microbiology and potential applications of aerobic methane oxidation coupled to denitrification(AME-D)process:A review[J]. Water Research, 2016, 90: 203-215.
(  0) 0) |
| [47] |
He Z F, Geng S, Cai C Y, et al. Anaerobic oxidation of methane coupled to nitrite reduction by halophilic marine NC10 bacteria[J]. Applied and Environmental Microbiology, 2015, 81(16): 5538-5545.
(  0) 0) |
| [48] |
Zhu G B, Zhou L L, Wang Y, et al. Biogeographical distribution of denitrifying anaerobic methane oxidizing bacteria in Chinese wetland ecosystems[J]. Environmental Microbiology Reports, 2015, 7(1): 128-138.
(  0) 0) |
| [49] |
Beal E J, House C H, Orphan V J. Manganese and iron-dependent marine methane oxidation[J]. Science, 2009, 325(5937): 184-187.
(  0) 0) |
| [50] |
Chang Y H, Cheng T W, Lai W J, et al. Microbial methane cycling in a terrestrial mud volcano in eastern Taiwan[J]. Environmental Microbiology, 2012, 14(4): 895-908.
(  0) 0) |
| [51] |
Scheller S, Yu H, Chadwick G L, et al. Artificial electron acceptors decouple archaeal methane oxidation from sulfate reduction[J]. Science, 2016, 351(6274): 703-707.
(  0) 0) |
| [52] |
Fu L, Li S W, Ding Z W, et al. Iron reduction in the DAMO/Shewanella oneidensis MR-1 coculture system and the fate of Fe(Ⅱ)[J]. Water Research, 2016, 88: 808-815.
(  0) 0) |
| [53] |
Ettwig K F, Zhu B L, Speth D, et al. Archaea catalyze iron-dependent anaerobic oxidation of methane[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(45): 12792-12796.
(  0) 0) |
| [54] |
He Z F, Zhang Q Y, Feng Y D, et al. Microbiological and environmental significance of metal-dependent anaerobic oxidation of methane[J]. Science of the Total Environment, 2018, 610/611: 759-768.
(  0) 0) |
| [55] |
DiChristina T J, Arnold R G, Lidstrom M E, et al. Dissimilative iron reduction by the marine Eubacterium Alteromonas putrefaciens strain 200[J]. Water Science and Technology, 1988, 20(11/12): 69-79.
(  0) 0) |
| [56] |
Lovley D R, Phillips E J P. Novel mode of microbial energy metabolism:Organic carbon oxidation coupled to dissimilatory reduction of iron or manganese[J]. Applied and Environmental Microbiology, 1988, 54(6): 1472-1480.
(  0) 0) |
| [57] |
Achtnich C, Bak F, Conrad R. Competition for electron donors among nitrate reducers, ferric iron reducers, sulfate reducers, and methanogens in anoxic paddy soil[J]. Biology and Fertility of Soils, 1995, 19(1): 65-72.
(  0) 0) |
| [58] |
Zeng C S, Wang W Q, Tong C. Effects of different exogenous electron acceptors and salt import on methane production potential of estuarine marsh soil (In Chinese)[J]. Geographical Research, 2008, 27(6): 1321-1330. [曾从盛, 王维奇, 仝川. 不同电子受体及盐分输入对河口湿地土壤甲烷产生潜力的影响[J]. 地理研究, 2008, 27(6): 1321-1330.]
(  0) 0) |
| [59] |
Min H, Tan Y L, Wu W X, et al. Isolation, purification and characterization of an anaerobic methane oxidizer (In Chinese)[J]. Journal of Zhejiang University(Agriculture and Life Sciences), 2002, 28(6): 619-624. [闵航, 谭玉龙, 吴伟祥, 等. 一个厌氧甲烷氧化菌菌株的分离、纯化和特征研究[J]. 浙江大学学报(农业与生命科学版), 2002, 28(6): 619-624.]
(  0) 0) |
| [60] |
Fu L, Li S W, Ding Z W, et al. Iron reduction in the DAMO/Shewanella oneidensis MR-1 coculture system and the fate of Fe(Ⅱ)[J]. Water Research, 2016, 88: 808-815.
(  0) 0) |
| [61] |
Egger M, Rasigraf O, Sapart C J, et al. Iron-mediated anaerobic oxidation of methane in brackish coastal sediments[J]. Environmental Science & Technology, 2015, 49(1): 277-283.
(  0) 0) |
| [62] |
Lu Y Z, Fu L, Ding J, et al. Cr(Ⅵ)reduction coupled with anaerobic oxidation of methane in a laboratory reactor[J]. Water Research, 2016, 102: 445-452.
(  0) 0) |
| [63] |
Chen Q F, Ma J J, Liu J H, et al. Characteristics of greenhouse gas emission in the Yellow River Delta wetland[J]. International Biodeterioration & Biodegradation, 2013, 85: 646-651.
(  0) 0) |
| [64] |
Chen Q F, Ma J J, Zhao C S, et al. The spatial and temporal variation characteristics of CH4 and CO2 emission flux under different land use types in the Yellow River Delta wetland[J]. Journal of Geoscience and Environment Protection, 2015, 3(6): 26-32.
(  0) 0) |
| [65] |
Milucka J, Ferdelman T G, Polerecky L, et al. Zero-valent sulphur is a key intermediate in marine methane oxidation[J]. Nature, 2012, 491(7425): 541-546.
(  0) 0) |
| [66] |
Lovley D R, Stolz J F, Nord G L Jr, et al. Anaerobic production of magnetite by a dissimilatory iron-reducing microorganism[J]. Nature, 1987, 330(6145): 252-254.
(  0) 0) |
| [67] |
Summers Z M, Fogarty H E, Leang C, et al. Direct exchange of electrons within aggregates of an evolved syntrophic coculture of anaerobic bacteria[J]. Science, 2010, 330(6009): 1413-1415.
(  0) 0) |
| [68] |
Rotaru A E, Thamdrup B. A new diet for methane oxidizers[J]. Science, 2016, 351(6274): 658.
(  0) 0) |
| [69] |
Wegener G, Krukenberg V, Riedel D, et al. Intercellular wiring enables Electron transfer between methanotrophic archaea and bacteria[J]. Nature, 2015, 526(7574): 587-590.
(  0) 0) |
| [70] |
McGlynn S E, Chadwick G L, Kempes C P, et al. Single cell activity reveals direct Electron transfer in methanotrophic consortia[J]. Nature, 2015, 526(7574): 531-535.
(  0) 0) |
| [71] |
Handelsman J, Rondon M R, Brady S F, et al. Molecular biological access to the chemistry of unknown soil microbes:A new frontier for natural products[J]. Chemistry & Biology, 1998, 5(10): R245-R249.
(  0) 0) |
| [72] |
Rodr|íguez-Valera F. Environmental genomics, the big picture?[J]. FEMS Microbiology Letters, 2004, 231(2): 153-158.
(  0) 0) |
| [73] |
Wongnate T, Sliwa D, Ginovska B, et al. The radical mechanism of biological methane synthesis by methyl-coenzyme M reductase[J]. Science, 2016, 352(6288): 953-958.
(  0) 0) |
| [74] |
Boxer S G, Kraft M L, Weber P K. Advances in imaging secondary ion mass spectrometry for biological samples[J]. Annual Review of Biophysics, 2009, 38(1): 53-74.
(  0) 0) |
 2020, Vol. 57
2020, Vol. 57